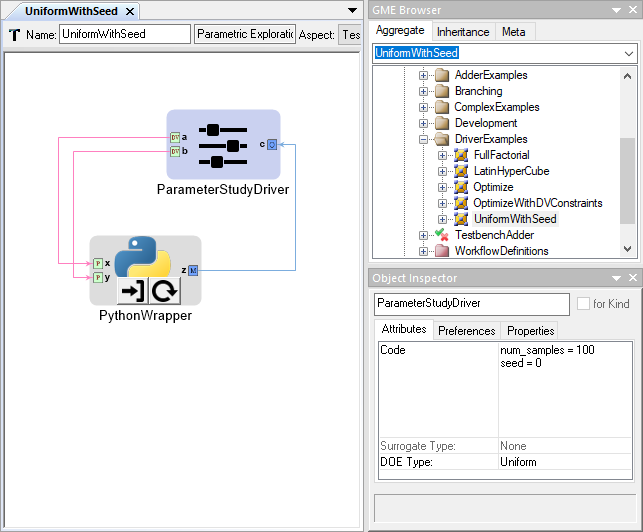
Parameter Study PET Driver¶
The Parameter Study Driver generates a set of inputs based on the type of Design of Experiments (DOE) that is being conducted. Then the driver evaluates the analysis workflow at each of the generated input sets and records the values of the objectives.
The Parameter Study is configured by setting and adjusting a number of its attributes. These attributes can be accessed by left-clicking a Parameter Study and then looking under the Attributes tab of the Object Inspector window.
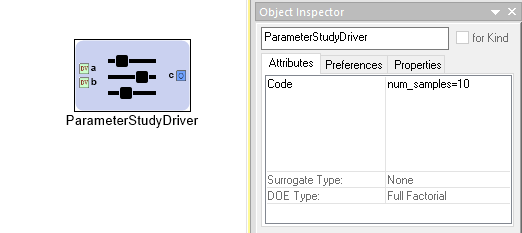
A Parameter Study Driver’s Attributes
Code¶
The Code attribute is used to pass configuration variables
to underlying MDAO engine. For example, in the image above you can
see we assigned the variable num_samples the value of 10.
You will use this field to set other values such as seed and num_levels.
See the DOE Types section below for more
information on what code variables need to be set for each type.
Surrogate Type¶
The Surrogate Type attribute is not currently documented.
DOE Types¶
The DOE Type attribute determines the sampling method by which the Parameter Study explores the Design Variable space. Different DOE Types can be selected by left-clicking the DOE Type attribute field and selecting the desired method. The different types and their accompanying configurations are described below.
Full Factorial¶
The Full Factorial DOE Type makes it easy to exhaustively explore a design space. This type generates dv^num_levels input cases where dv is the number of Design Variables (or factors) in our driver and num_levels, set in the Code attribute, is the number of discrete values to consider from the range specified for each design variable.
If num_levels = 1, then each Design Variable will have one level equal to the lower bound specified in the Range Attribute of the Design Variable. In the case of a single value defined in the Range attribute, the level is simply the value. If num_levels = 2, then each Design Variable will have two levels equal to the lower and upper bounds specified in the range attribute of the Design Variable. If num_levels = 3 or greater, then each Design Variable will have levels evenly distributed across its range, including the lower and upper bounds.
For example, if the PET has two Design Variables x and y, both with a Range of -10,10 and the Code attribute defined with num_levels = 3, then the following combination of inputs would be simulated:
| row | x | y |
|---|---|---|
| 1 | -10 | -10 |
| 2 | -10 | 0 |
| 3 | -10 | 10 |
| 4 | 0 | -10 |
| 5 | 0 | 0 |
| 6 | 0 | 10 |
| 7 | 10 | -10 |
| 8 | 10 | 0 |
| 9 | 10 | 10 |
Central Composite¶
This DOE type is currently unsupported.
Opt Latin Hypercube¶
The Opt[imized] Latin Hypercube type is a pre-determined samples driver that seeks to produce good coverage across all the dimensions (or Design Variables). This is preferred to a Uniform type of sampling in most cases as you have a higher probability of an evenly-sampled independent variables space.
This pre-ddetermine samples driver is implemented using an evolutionary algorithm to optimize the Morris-Mitchell sampling criterion. [1] This type of DOE supports a number of configuration options to assign in the code block:
- num_samples - This specifies the total number of samples to generate. It must be greater than or equal to 2.
- seed - This is used to seed the pseudorandom number generator (PRNG). If it is assigned, the generated Latin Hypercube Design (or input cases) will always be the same when this PET is run, i.e. the behavior will be deterministic. (Default: random)
- generations - This determines how many generations the genetic algorithm will evolve over. (Default: 2)
- population - This sets the size of the population used in the evolutionary optimization. (Default: 20)
- norm-method - This determines the method for determining the vector norm. 1-norm is faster, but less accurate. (Choices: [“1-norm”, “2-norm”], Default: “1-norm”)
Uniform¶
The Uniform type generates a input cases by simply sampling each Design Variable using a uniform or discrete uniform distribution across the interval defined in its Range attribute. It will generate as many input cases as are specified by the num_samples assignment in the Code attribute.
As with the Opt Latin Hypercube DOE type, you can realize deterministic behavior by setting the seed parameter in the code block. Take as an example one of the PETs in the analysis-blocks project of the openmeta-examples-and-templates repository:
Since the seed is set to 0, the results will always be the same when this PET is executed. Try it yourself and see if your results match!

CSV File¶
The CSV File type allows for an arbitrary set of test cases to be specified in a CSV file and then executed with the given analysis workflow. This is useful when you have a number of edge cases you need to test.
The input file is selected by placing the path, relative to the project
directory (i.e. the location of the current .mga file), in a
filename='<path>' assignment in the Code attribute of the
Parameter Study Driver. This file will be copied to the execution
directory when the PET is executed.
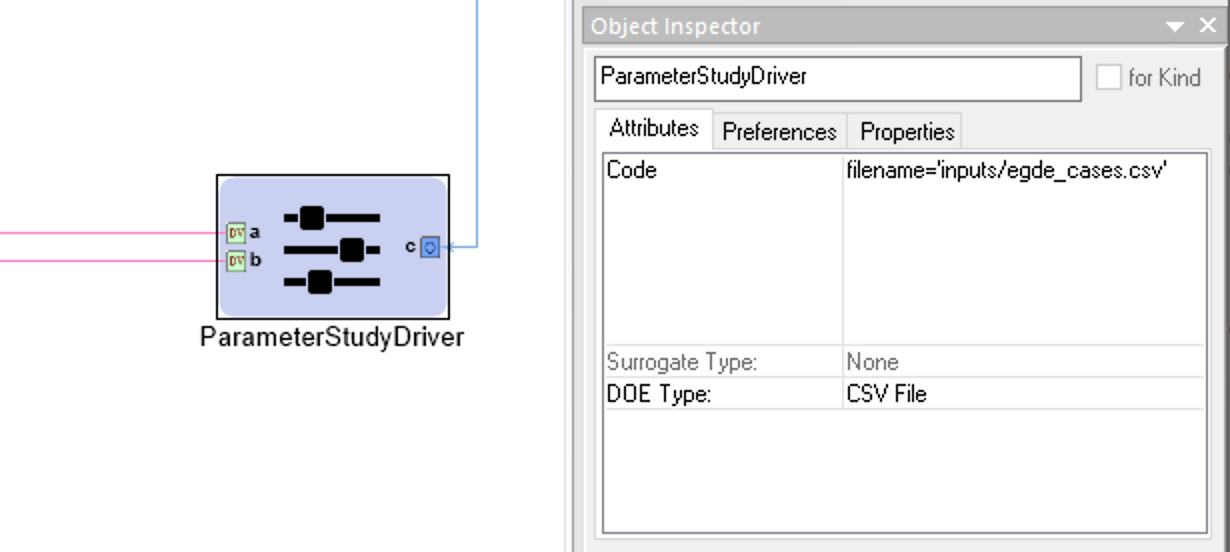
Example CSV File DOE Type Configuration for a Parameter Study Driver
All design variables that are unrepresented in the input CSV file will be assigned a value that is the average of the interval specified in that design variable’s Range attribute in the Parameter Study Driver. Extra columns that don’t match any of the design variables are allowed in the input CSV, but they are ignored.
Footnotes
| [1] | http://openmdao.org/releases/0.1.4/docs/srcdocs/packages/openmdao.lib.html#openmdao-lib-doegenerators-optlh-py |